January 27, 2026
by Mikhail Elyashberg, Leading Researcher, ACD/Labs
Computer-Assisted Elucidation of Verrucosin C Using Structure Elucidator Suite
Fenical and co-workers [1] investigated the Verrucosispora sp. strain TAA-831, a member of the Micromonosporaceae family, collected in the Northern Marianas Islands. A multitude of unknown natural products were found after chemical profiling of this strain. This was confirmed via MS and NMR-based dereplication techniques. The authors undertook a comprehensive chemical analysis to structurally characterize these assumed novel metabolites which resulted in the isolation of five new polyketides, the verrucosins A−E. Their structures were determined through a synergistic combination of spectroscopic, computational, and genomic approaches.
The spectroscopic data used by the authors for structure elucidation of verrucosin C (1) were used by us to challenge ACD/Structure Elucidator.
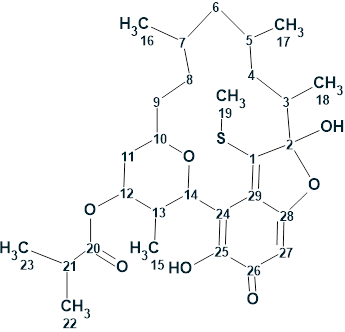
1
Verrucosin C was isolated as a bright orange oil. It exhibited a deprotonated molecule in the (−)-HRESIMS at m/z 533.2585, consistent with the molecular formula C29H41O7S− (calcd for 533.2574) and nine degrees of unsaturation. The 13C, 1H, HMBC and COSY spectroscopic data are presented in Table 1.
Table 1. NMR spectroscopic data of verrucosin C [1].
| C/X Label | δC | δCcalc HOSE | XHn | δH | COSY | H to C HMBC |
| C 1 | 154.1 | 155.03 | C | |||
| C 2 | 117.8 | 119.85 | C | |||
| C 3 | 41.5 | 41.58 | CH | 1.98 | 1.23, 1.29 | C 18, C 5, C 4, C 2 |
| C 4 | 36.9 | 38.82 | CH2 | 1.29 | 1.37, 1.98 | C 18, C 2 |
| C 4 | 36.9 | 38.82 | CH2 | 0.49 | C 18, C 17, C 5, C 3 | |
| C 5 | 33.5 | 29.32 | CH | 1.37 | 0.88, 0.93, 1.29 | |
| C 6 | 47.8 | 44.82 | CH2 | 0.88 | 1.37, 1.76 | C 4 |
| C 6 | 47.8 | 44.82 | CH2 | 0.81 | C 5 | |
| C 7 | 27.2 | 30.61 | CH | 1.76 | 0.79, 0.88, 1.40 | |
| C 8 | 32.5 | 33.95 | CH2 | 1.4 | 1.31, 1.76 | |
| C 8 | 32.5 | 33.95 | CH2 | 0.89 | ||
| C 9 | 30.9 | 33.49 | CH2 | 1.41 | ||
| C 9 | 30.9 | 33.49 | CH2 | 1.31 | 1.40, 3.88 | C 8, C 10 |
| C 10 | 69.3 | 73.04 | CH | 3.88 | 1.31, 1.51 | |
| C 11 | 37.6 | 34.41 | CH2 | 1.72 | C 10 | |
| C 11 | 37.6 | 34.41 | CH2 | 1.51 | 3.88, 5.06 | C 12 |
| C 12 | 71.5 | 71.67 | CH | 5.06 | 1.51, 2.84 | C 10, C 14, C 20 |
| C 13 | 35.7 | 33.9 | CH | 2.84 | 0.78, 5.06, 5.39 | C 15, C 14 |
| C 14 | 74.2 | 76.81 | CH | 5.39 | 2.84 | C 13, C 10, C 12, C 24, C 29, C 25 |
| C 15 | 13.2 | 11.25 | CH3 | 0.78 | 2.84 | C 13, C 12, C 14 |
| C 16 | 18.8 | 20.44 | CH3 | 0.79 | 1.76 | C 7, C 8, C 6 |
| C 17 | 23.7 | 20.41 | CH3 | 0.93 | 1.37 | C 5, C 4, C 6 |
| C 18 | 15.6 | 12.28 | CH3 | 1.23 | 1.98 | C 4, C 3, C 2 |
| C 19 | 15.2 | 17.5 | CH3 | 2.71 | C 1 | |
| C 20 | 175.3 | 178.51 | C | |||
| C 21 | 34 | 34.99 | CH | 2.6 | 1.17 | C 22, C 23, C 20 |
| C 22 | 18.8 | 19.6 | CH3 | 1.17 | 2.6 | C 23, C 21, C 20 |
| C 23 | 18.9 | 19.6 | CH3 | 1.17 | ||
| C 24 | 109.5 | 113.87 | C | |||
| C 25 | 149.2 | 148.9 | C | |||
| C 26 | 180.4 | 182.75 | C | |||
| C 27 | 95 | 95.8 | CH | 5.7 | C 29, C 25, C 28, C 26 | |
| C 28 | 169.4 | 172.35 | C | |||
| C 29 | 125.7 | 130.95 | C | |||
| O 1 | OH | 8.04 | C 3, C 2, C 1 | |||
| O 2 | OH | 9.26 | C 24, C 25, C 26 |
These data were entered in ACD/Structure Elucidator, and a molecular connectivity diagram (MCD) was automatically created (Figure 1).
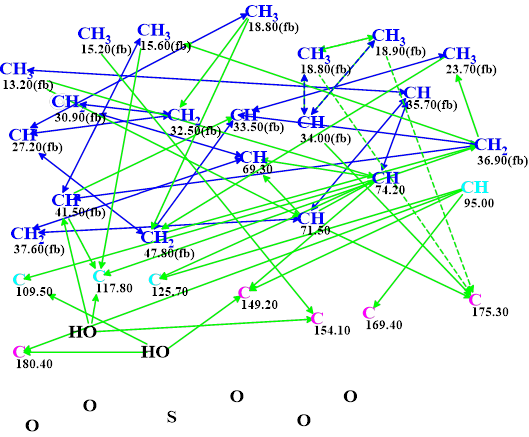
Figure 1. Molecular connectivity diagram (MCD) of verrucosin C. Hybridizations of carbon atoms are marked by corresponding colors: sp2 – violet, sp3 – blue, not sp – light blue. Labels “ob” and “fb” are set by the program to carbon atoms for which neighboring with an heteroatom is either obligatory or forbidden respectively. The HMBC connectivities are marked with green arrows, while COSY connectivities are marked with blue arrows. Ambiguous connectivities are marked with dotted lines.
All atom properties shown in the MCD were determined by the program automatically. Note that the four light blue carbon atoms at 95.00, 109.50, 117.80 and 125.7 ppm are characterized by an ambiguous hybridization not sp (i.e., either sp3 or sp2). Such a hybridization assignment considers the possibility of the O-C-O fragment presence in the structure. All possible hybridizations of the mentioned atoms will be tried during the structure generation process. No manual edits of the MCD were made.
The MCD checking for 2D NMR data consistency showed that no nonstandard correlations (nJCH,HH, n>3) were present in the spectra, so strict structure generation accompanied with 13C chemical shift prediction was initiated. Generation was completed with the following results: k = 1496 → (Structural Filtering) → 2, tg = 10s.
The two structures generated are shown in Figure 2.
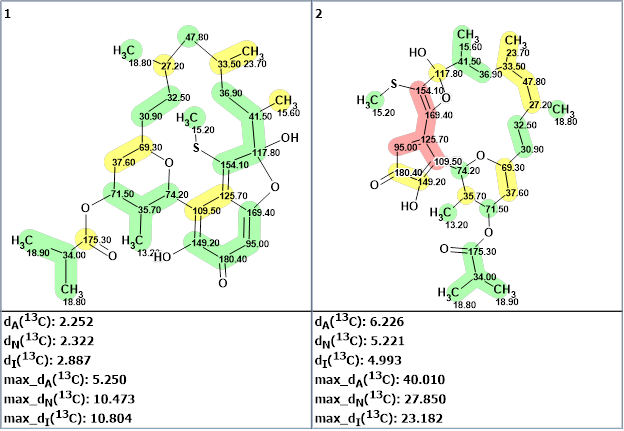
Figure 2. The output file for verrucosin C. 13C chemical shift prediction was carried out using the HOSE code-based method, the neural networks, and the incremental approach. Average deviations of 13C chemical shifts determined by these methods are denoted as dA, dN and dI respectively. Each atom is colored to mark a difference between its experimental and calculated 13C chemical shifts. Green represents a difference between 0 to 3 ppm, yellow is between 3 and 15 ppm and red is more than 15 ppm.
We see that the first structure in Figure 1 is verrucosin C, while the second structure should be rejected not only by the very large average and maximum deviations but also due to the presence of a 4-membered cycle which is quite unusual for natural products.
Therefore, the structure of a large molecule (C29H42O7S, m/z ~ 533) was quickly (in 10s) and unambiguously elucidated using Structure Elucidator in a fully automatic mode. This was possible as the 2D NMR spectroscopic data were consistent and complete enough in this case. The comprehensiveness of the initial data (a set of “axioms”) was possible by the large amount of hydrogen atoms in the molecular formula.
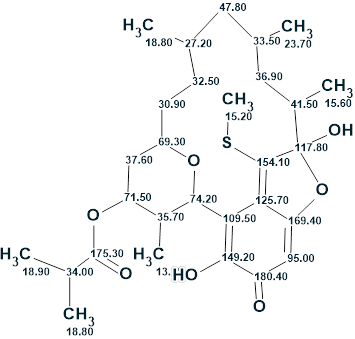
The structure of verrucosin C with assigned 13C chemical shifts is shown below:
References
- C. Holland, R. Iizumi, V.V. Shende, W. Fenical. (2025). Isolation of Verrucosins A−E from a Marine Verrucosispora sp. Reveals a Unifying Biosynthetic Hypothesis for Linear and Macrocyclic Polyketides. J. Nat. Prod., 88(7), 1631–1642. https://doi.org/10.1021/acs.jnatprod.5c00373


