February 26, 2026
by Mikhail Elyashberg, Leading Researcher, ACD/Labs
Computer-Assisted Elucidation of Biatractylonoid D Using Structure Elucidator Suite
Dimeric sesquiterpenoids, characterized by a C30 carbon framework formed through diverse dimerization modes of two homologous or heterologous monomeric units, are found predominantly in plants of the Asteraceae family. Owing to their structurally complex architectures and notable biological activities, these compounds have attracted considerable attention from organic chemists. To date, 291 disesquiterpenoids have been reported from Asteraceae species.
In contrast, dimers derived from eudesmane-type sesquiterpenoids are comparatively uncommon. Atractylodes macrocephala Koidz., a perennial herb of the Asteraceae family widely distributed in China, Korea, and Japan, is a well-known medicinal plant in traditional Chinese medicine, where its rhizomes are extensively used. Thus far, only six eudesmane-type sesquiterpenoid dimers have been isolated from the rhizomes of A. macrocephala.
Zhang et al. [1] recently and unexpectedly identified five eudesmane-type dimeric sesquiterpenoids with two previously unknown carbon skeletons, designated biatractylonoids A–E. These compounds possess a novel rearranged eudesmane dimer framework incorporating a distinctive 2-oxaspiro[4.4]onane motif. Notably, biatractylonoid D (1) exhibits an unprecedented heptacyclic ring system with a unique 6/6/5/5/5/5/6 ring fusion pattern, in which two monomeric units are linked through an unusual dimerization pathway involving a five-membered oxygen-containing heterocycle.
Biatractylonoid D (1) was obtained as a white powder. It had a molecular formula of C30H38O5 from HR-ESI-MS: m/z 479.2788 [M + H]+ (calcd. for C30H39O5 479.2792). The structure elucidation of this compound was carried out using 1D and 2D NMR (HSQC, HMBC and COSY) data in combination with comparison with spectra of similar compounds (biatractylonoids A-C) identified earlier in [1].
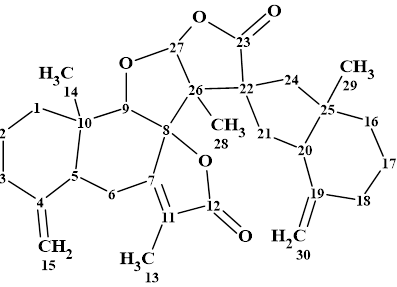
1
The molecular formula and spectroscopic data (Table 1) obtained for compound 1 were used for challenging ACD/Structure Elucidator
Table 1. 1D and 2D NMR spectroscopic data of biatractylonoid D [1].
| Label | δC | δC clc (HOSE) | XHn | δH | 1H Mul. | COSY | H to C HMBC |
| C 1 | 35.6 | 35.75 | CH2 | 2.05 | u | 1.51 | |
| C 1 | 35.6 | 35.75 | CH2 | 1.36 | u | ||
| C 2 | 23.1 | 23.57 | CH2 | 1.6 | u | ||
| C 2 | 23.1 | 23.57 | CH2 | 1.51 | u | 2.01, 2.05 | |
| C 3 | 37.6 | 36.17 | CH2 | 2.41 | u | ||
| C 3 | 37.6 | 36.17 | CH2 | 2.01 | u | 1.51 | C 15 |
| C 4 | 149.2 | 150.44 | C | ||||
| C 5 | 40.3 | 43.15 | CH | 2.74 | u | 2.89 | C 15 |
| C 6 | 27.7 | 26.24 | CH2 | 2.71 | u | ||
| C 6 | 27.7 | 26.24 | CH2 | 2.89 | u | 2.74 | C 10, C 8 |
| C 7 | 163.7 | 161.99 | C | ||||
| C 8 | 94.8 | 95.09 | C | ||||
| C 9 | 92.7 | 86.88 | CH | 4.09 | s | ||
| C 10 | 40.7 | 38.98 | C | ||||
| C 11 | 127.6 | 124.18 | C | ||||
| C 12 | 173.5 | 173.31 | C | ||||
| C 13 | 8.8 | 8.28 | CH3 | 1.83 | s | C 7, C 12 | |
| C 14 | 16.9 | 23.74 | CH3 | 0.52 | s | C 1, C 9 | |
| C 15 | 109.1 | 107.2 | CH2 | 4.99 | s | ||
| C 15 | 109.1 | 107.2 | CH2 | 4.86 | s | ||
| C 16 | 40.2 | 39.78 | CH2 | 1.69 | u | 1.64 | |
| C 16 | 40.2 | 39.78 | CH2 | 1.25 | u | ||
| C 17 | 24.4 | 23.08 | CH2 | 1.64 | u | 1.69, 2.27 | |
| C 18 | 35.7 | 35.07 | CH2 | 2.27 | u | 1.64 | C 30 |
| C 18 | 35.7 | 35.07 | CH2 | 1.94 | u | ||
| C 19 | 148.8 | 148.87 | C | ||||
| C 20 | 54.1 | 53.26 | CH | 1.97 | u | 1.8 | C 30 |
| C 21 | 36.8 | 33.55 | CH2 | 1.8 | u | 1.97 | C 25, C 26, C 23 |
| C 21 | 36.8 | 33.55 | CH2 | 2.16 | u | ||
| C 22 | 57.6 | 55.08 | C | ||||
| C 23 | 181.1 | 176.82 | C | ||||
| C 24 | 44.3 | 42.79 | CH2 | 2.23 | u | C 23 | |
| C 24 | 44.3 | 42.79 | CH2 | 1.09 | u | ||
| C 25 | 44.1 | 43.35 | C | ||||
| C 26 | 62.3 | 55.21 | C | ||||
| C 27 | 110.1 | 110.3 | CH | 5.88 | s | C 28, C 8, C 23 | |
| C 28 | 13.5 | 15.43 | CH3 | 1.33 | s | C 22, C 8 | |
| C 29 | 18.7 | 26.27 | CH3 | 0.87 | s | C 16, C 24 | |
| C 30 | 106.2 | 106.35 | CH2 | 4.42 | s | ||
| C 30 | 106.2 | 106.35 | CH2 | 4.75 | s |
Valuable preliminary information about the functional groups included in the structure can be extracted from the IR spectrum, shown in Figure 1.
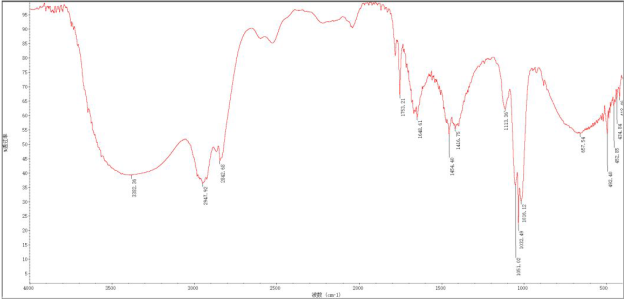
Figure 1. IR spectrum of biatractylonoid D presented in article [1]
It is evident that the sample, unfortunately, contained large amounts of water (bands 3400, 1650 and 660 cm-1) and, consequently, could not be used for drawing conclusions regarding the presence or absence of hydroxyl groups in the molecule. However, the absorption band at 1753 cm-1 implies the presence of 5-membered lactone.
The data was entered into the program from Table 1, and a Molecular Connectivity Diagram, MCD, was created (Figure 2).
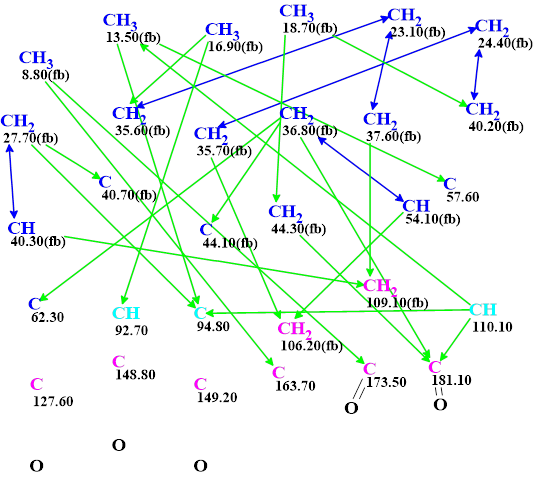
Figure 2. Molecular connectivity diagram (MCD) of biatractylonoid D. The hybridizations of carbon atoms are marked by the corresponding colors: sp2 – violet, sp3 – blue, not sp – light blue. The label “fb” is set by the program to carbon atoms for which neighboring with a heteroatom is forbidden (fb). HMBC connectivities are marked by green arrows, while COSY connectivities by blue arrows. The evident C=O bonds were drawn manually to accelerate structure generation
Visual analysis of the MCD allows us to conclude that the CASE-based structure elucidation of this compound will be time consuming. Indeed, we see that three carbon atoms C 127.60, 148.8 and 149.2 have no correlation in HMBC and COSY spectra. This means that the program will exhaust all possibilities of inserting these carbons into structures during the generation process. Another peculiarity of the MCD is the presence of three light blue carbons C 92.7, C 94.8 and C 110 with ambiguous hybridization – sp2 or sp3. All possible combinations of hybridization must also be tried during the structure generation to accommodate for this. These combinations are shown below:
| C 97.2 | C 94.8 | C 110.10 |
| sp3 | sp3 | sp3 |
| sp3 | sp2 | sp2 |
| sp3 | sp3 | sp2 |
| sp2 | sp2 | sp2 |
| sp2 | sp3 | sp3 |
| sp2 | sp2 | sp3 |
As the generation time was expected to be long this procedure was initiated in the evening, accepting that it will be completed overnight. Structure generation was accompanied by 13C chemical shift prediction and structure filtering. The structures for which the average deviations of predicted chemical shifts from experimental ones exceeded 5 ppm were rejected. Results: k = 235,040 → (Structure Filtering) → 1231 → (Duplicate Removal) → 337, tg = 12 h 15 m. Even though the structure generation time was more than twelve hours, this procedure was performed autonomously without requiring human input. The structures in the output file were ranked in increasing order of average deviations. The twelve top ranked structures are shown in Figure 3.
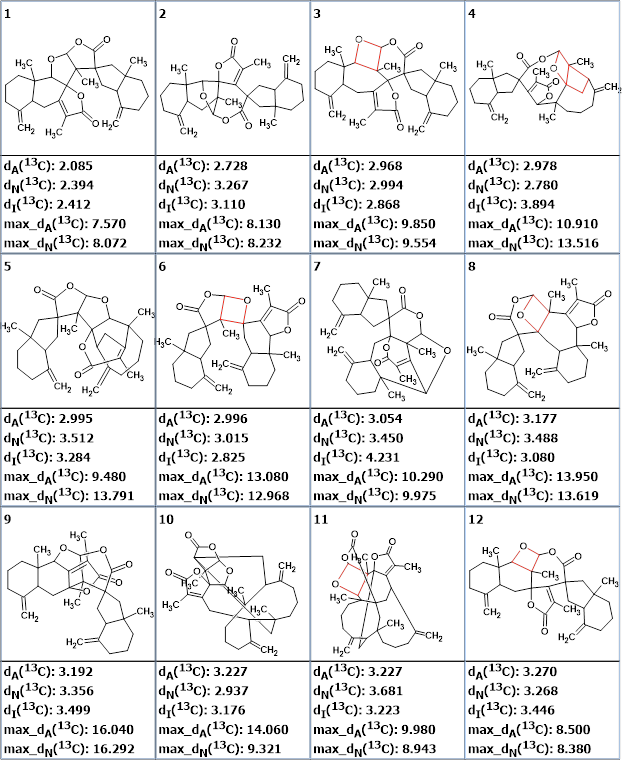
Figure 3. The structures in the ranked output file. 13C chemical shift prediction was carried out using the HOSE code-based method, the neural networks, and the incremental approach. Average deviations of 13C chemical shifts determined by these methods are denoted as dA, dN and dI correspondingly. Oxetanes are marked by red color in structures.
We see that the best structure is identical to that which was determined in [1] and confirmed by DFT calculations and by X-Ray analysis. The figure shows that half of the structures contain an oxetane ring (marked in red) which is rarely observed in natural products. With this in mind, we removed the structures containing an oxetane ring from the output file, so 112 structures out of the 337 remained. The top eight structures of the newly ranked file are presented in Figure 4.
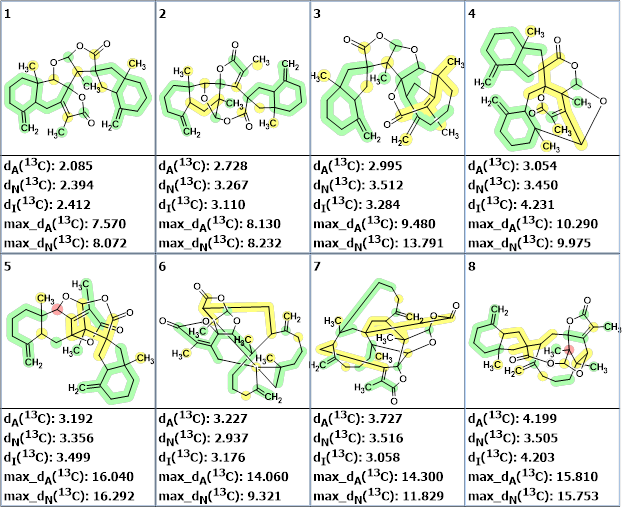
Figure 4. The eight top ranked structures of the reduced output file. 13C chemical shift prediction was carried out using the HOSE code-based method, the neural networks, and the incremental approach. Average deviations of 13C chemical shifts determined by these methods are denoted as dA, dN and dI correspondingly. Each atom is colored to mark a difference between its experimental and calculated 13C chemical shifts. The green color represents a difference between 0 and 3 ppm, yellow was between 3 and 15 ppm and red was more than 15 ppm.
Furthermore, we calculated the DP4 probabilities for the first four structures shown in Figure 4, and the results are presented in Figure 5.
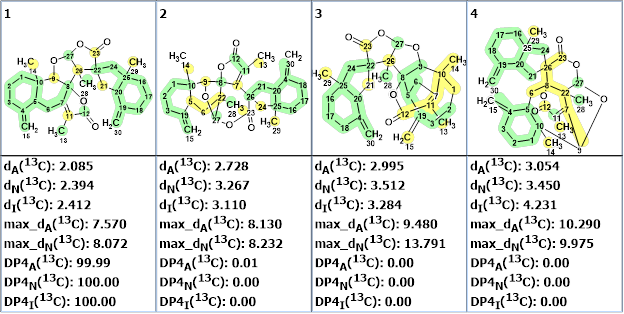

Figure 5. The DP4A, DP4N and DP4I probabilities of structure correctness calculated by the program.
The calculations confirmed the validity of the first ranked structure for which the DP4 probabilities were determined by all three prediction methods to be equal to 100%.
Furthermore, it seemed interesting to us from a methodological point of view to evaluate how atoms having no 2D NMR correlations and light blue atoms with undefined hybridization separately affect the generation time. For this task, we set sp3 hybridization for atoms C 92.7, C 94.8 and C 110 ppm and supplied them with label “ob” (neighborhood with a heteroatom is obligatory), and then structure generation was repeated. It gave the following results: k = 63,572 → (Structure Filtering) → 417 → (Duplicate Removal) → 337, tg = 4h 35m
Generating structures is a complex process, but this computational experiment suggests that most of the time was spent trying out all possible combinations of hybridizations of the three light blue carbon atoms. When the hybridizations of these atoms were specified, the time was reduced from 12h 15m down to 4h 35m.
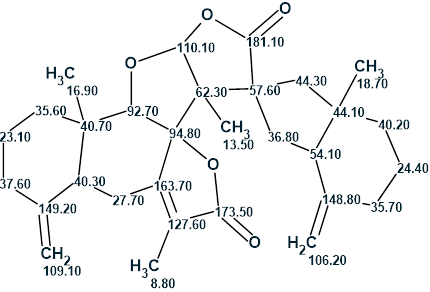
Thus, the complex structure of biatractylonoid D characterized by an unprecedented skeleton was unambiguously elucidated by Structure Elucidator in an automatic mode after a structure generation performed over 12 hours. The obtained structure together with the 13C chemical shift assignment is shown below:
References
- Zhang, L. Yin, S. Yu, H. Jin, M. Yu, L. Li, T. Zhang, K. Zeng, Z. Zou. (2025). Biatractylonoids A−E, Five Eudesmane-Type Sesquiterpenoid Dimers from the Rhizomes of Atractylodes macrocephala with Antiinflammatory Activity. J. Org. Chem., 90(44), 15748−15759. https://doi.org/10.1021/acs.joc.5c01988


