January 1, 2014
by Mikhail Elyashberg, Leading Researcher, ACD/Labs
Epohelmin A
In an earlier review article [1], a wide variety of examples of structure revision utilizing ACD/Structure Elucidator Suite were described. This month we re-visit one of them which demonstrates how the software allows a researcher to avoid complex multi-stage synthesis to refute a wrong proposed structure and prove the revised one.
Sakano et al [2] reported the isolation of the novel lanosterol synthase inhibitors epohelmins A (1) and B (2). The structures were determined by detailed spectroscopic analysis (including 2D NMR spectra) and proposed to be novel 9-oxa-4-azabicyclo[6.1.0]-nonanes. However these structure assignments raised doubts based on both chemical and spectroscopic grounds [3].
Snider and Gao [3] comprehensively analyzed both the spectral and chemical aspects of the study of epohelmins A and B and suggested structures 3 and 4 correspondingly as being more appropriate hypotheses:
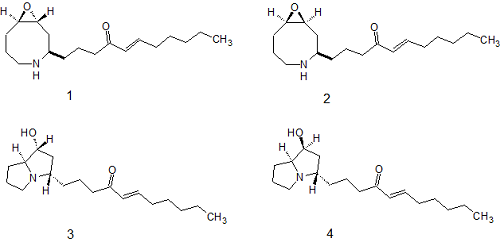
To validate their suggestions, the authors [3] developed an eight step synthesis of epohelmin A (1) and an 11 step synthesis of epohelmin B (2). Preparation of epohelmin A (1) and epohelmin B (2) is illustrated by Scheme 1.
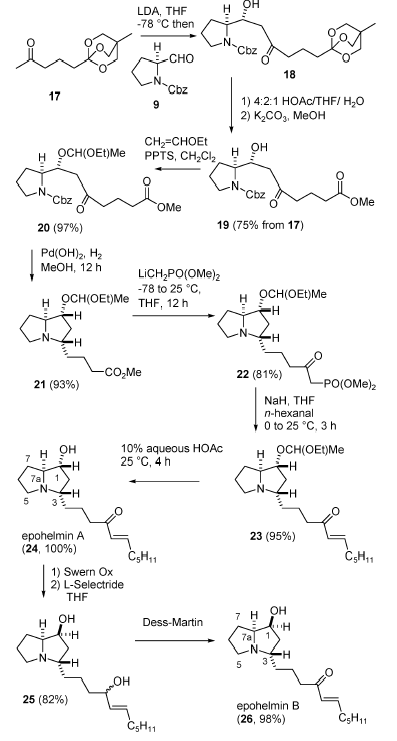
Scheme 1: The synthesis method for epohelmin A (1) and epohelmin B (2).
The 1H and 13C NMR spectra of 3 and 4 were identical to those reported for epohelmin A (1) and epohelmin B (2), and the revised structures of these compounds were therefore unambiguously established via chemical synthesis.
The 2D NMR spectra of the investigated compounds were not available for us, so it was only possible to predict and compare the 13C NMR spectra of competing structures 1 and 3, along with a review of the discrepancies between the predicted and experimental data (see Table 1).
| Structure | d(HOSE), ppm | d(NN), ppm | d(max) | R2(HOSE) | R2(NN) |
| 1 | 4.00 | 4.17 | 21.4 | 0.978 | 0.980 |
| 3 | 1.23 | 1.25 | 4.84 | 0.999 | 0.999 |
Table 1 unambiguously shows that structure 3 is superior over structure 1. It is likely that if 2D NMR data were available to the researchers then application of StrucEluc would deliver the correct structure very quickly and structure 1 would immediately be rejected by the program due to the very large deviations, especially with a d(max) value of 21.4 ppm. Multi-step syntheses would also not be necessary to resolve the structural problem. However, at the same time the method of synthesizing epohelmin A and epohelmin B would not be developed! This contradictory peculiarity of structural revision work was emphasized strongly in a subsequent review article [4] where a number of striking examples were given.
However, let us demonstrate how Structure Elucidator Suite would allow a researcher to not only reject the erroneous structure from the very beginning, but to easily find the correct one even if the 2D NMR spectra are not available.
The published 13C NMR spectral data from [2] was entered into Structure Elucidator, and the Molecular Connectivity Diagram (MCD) was created. The carbon atoms included into the side chains of both the original and revised structures (Figure 1) have precisely predicted 13C chemical shifts, therefore it is clear that the differences in calculated deviations for these structures must be due to differences in their cyclic fragments.
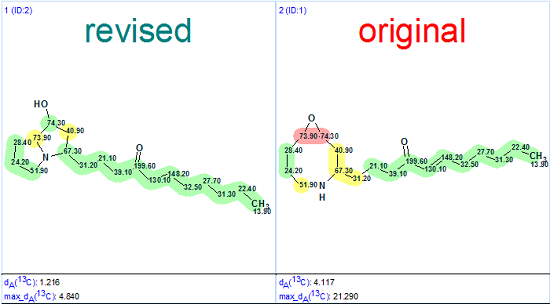
Figure 1.
With this in mind the atoms included into the side chain were connected by hand on the MCD to complete the common part of both structures (Figure 2).
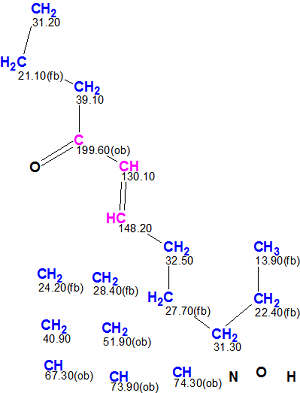
Figure 2. The Molecular Connectivity Diagram, including the manually modified hydrocarbon chain common to both revised and original structures.
The software was set to generate structures run from the MCD, which gave the following results: k = 10134 → 4180 → 167, tg = 25 s. Then 13C chemical shifts were predicted for the structures in the output file, and the structures were ranked by chemical shift deviations. The most interesting structures of the ranked file are presented in Figure 3.
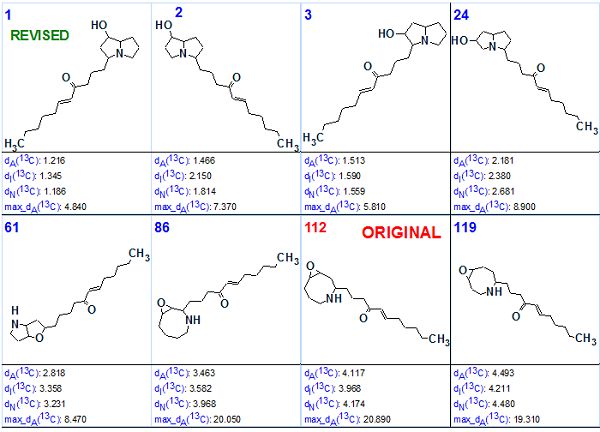
Figure 3. Portions of the ranked structure list following structure generation.
Figure 3 shows that the correct (revised) structure was selected as the most probable, and it is also distinguished among other isomers of the family (structures ranked as 2, 3, 24 and 61). Three isomers of the wrong structure family occupy 86, 112 and 119 positions, while structure 1 originally proposed by authors [2] is 112th (!) in the ranked file.
This example clearly demonstrates that even with only 13C NMR data, Structure Elucidator Suite can be used to refute incorrect structures and confirm the correct one. This method also avoids the need for time-consuming and costly multi-step synthesis to confirm the correct structure. Although it is obviously eventually beneficial to confirm a feasible synthesis pathway, this information is superfluous in the work of quickly obtaining the correct structure.
References
- M. Elyashberg, A. Williams, K.Blinov. Structural revisions of natural products by Computer Assisted Structure Elucidation (CASE) Systems. Nat. Prod. Rep., 27(9):1296–1328, 2010.
- Y. Sakano, M. Shibuya, Y. Yamaguchi, R. Masuma, H. Tomada, S. Omura, Y. Ebizuka, J. Antibiot. 57:564–568, 2004.
- B. B. Snider and X. Gao., Structure Revision and Syntheses of Epohelmins A and B. Org. Lett. 7:4419–4422, 2005.
- K.C Nicolaou; S.A. Snyder. Chasing Molecules That Were Never There: Misassigned Natural Products and the Role of Chemical Synthesis in Modern Structure Elucidation. Angew. Chem. Int. Ed. 44:1012–1044, 2005.


